The Molecular Information Systems Lab (MISL) at the University of Washington explores the intersection of information technology and molecular biology using in-silico and wet lab experiments. A partnership between UW Computer Science, Electrical Engineering, and Microsoft Research, MISL brings together faculty, students and research scientists with expertise in computer architecture, programming languages, synthetic biology, and biochemistry.
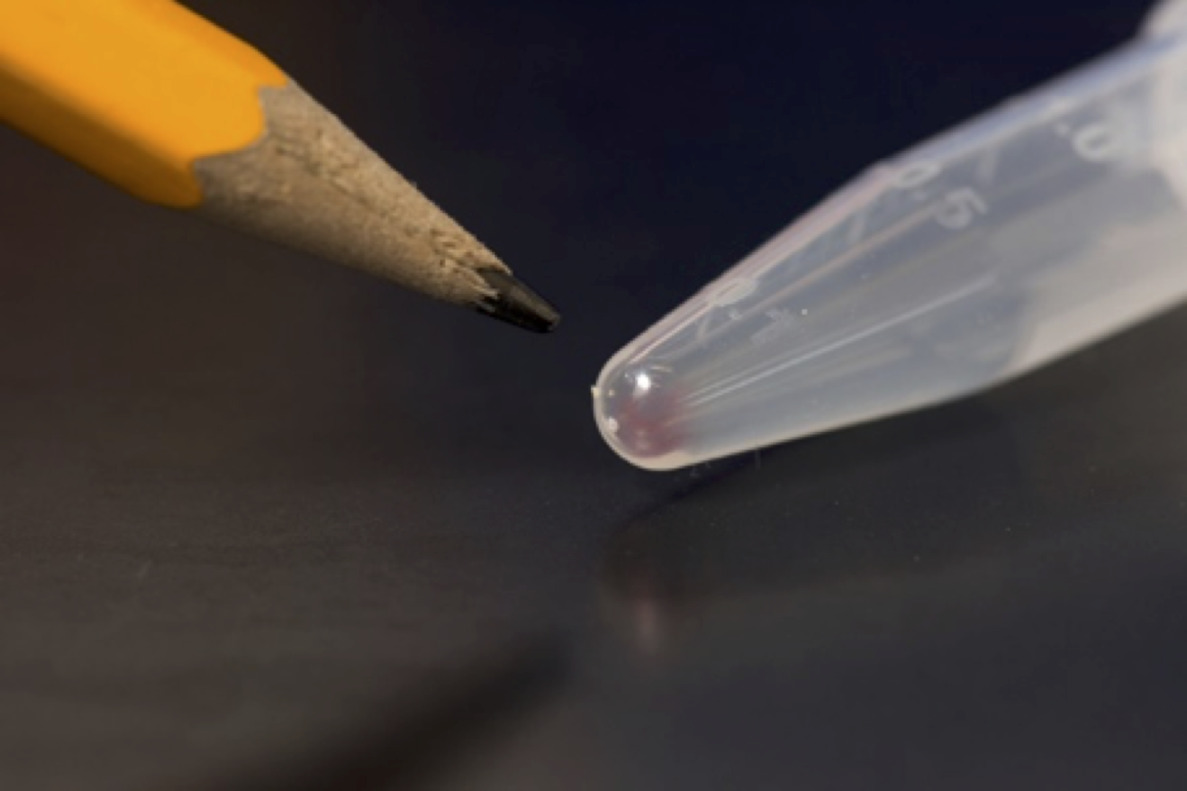

Why are we excited about molecular storage? The faint pink smear of DNA in the photo can hold over 10 terabytes of data. And it can last for a long time.
The integration of silicon and biomolecular systems is a promising direction of research that is motivated by potentially transformative outcomes. For example, molecular storage and computing components can be incredibly small, engineered with atomic precision, operate autonomously, and are capable of directly interfacing with natural biological systems for "near-data" processing tasks. However, for such systems to have broader real-world use, they must become faster, scalable, and more accessible to non-experts. Towards these goals, we apply molecular engineering, machine learning and nascent nanopore technology to overcome current limitations in molecular information processing systems.
Our Commitment
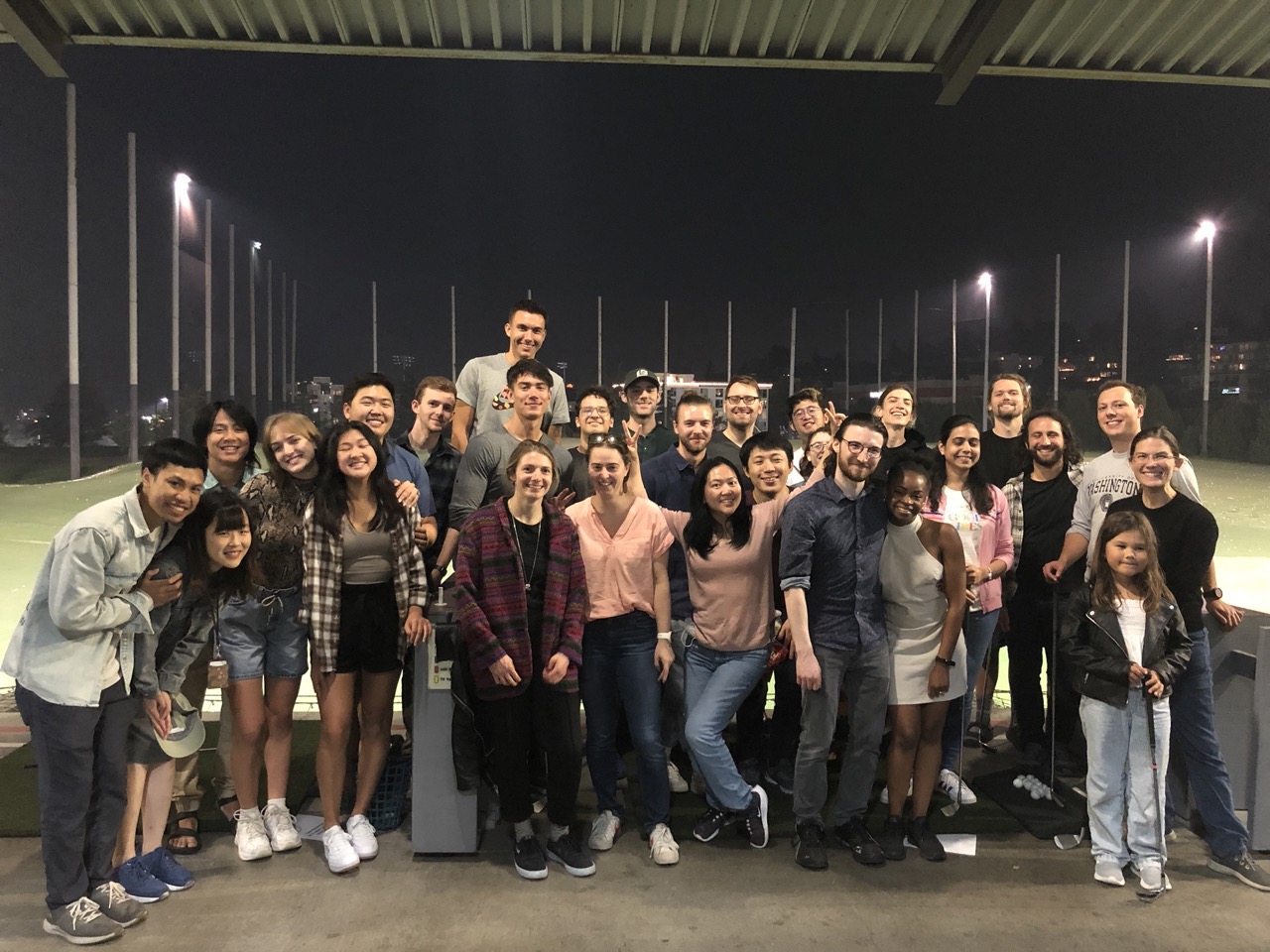

Our mission at the Molecular Information Systems Lab is to bring together researchers from diverse scientific backgrounds and life histories to explore a future where electronics and biology engineered together lead to new systems that make life on earth better. We believe that diverse perspectives and experiences are critical to our success as an organization, because a more diverse and inclusive team is more creative and effective. We strive to create a welcoming and engaging professional environment free of discrimination.
We, the Principal Investigators at MISL, commit to supporting people, especially those from under-represented groups, in developing their ideas and achieving their goals. We seek to connect people, especially those from under-represented groups, to opportunities that enable them to achieve their goals and fulfill their potential. We are dedicated to attracting and retaining a diverse group of individuals from all backgrounds and experiences, and to promoting an atmosphere of respect and open-mindedness. One way we stand by this commitment is through partnerships with local organizations that promote STEM education and real-world involvement at the early stages (e.g. K-12) to enhance equitable opportunity in the future.
Finally, our lab is committed to ongoing efforts to evaluate and address any barriers to DEI in our space, and to continuously improve in these pursuits. We look forward to any questions, thoughts, opportunities, or suggestions you may have for us, please feel free to reach out at any time.
-The MISL Leadership: Luis Ceze, Karin Strauss, Chris Takahashi, Jeff Nivala, Georg Seelig, and Chris Thachuk
News
-
May 19, 2026
-
May 13, 2026
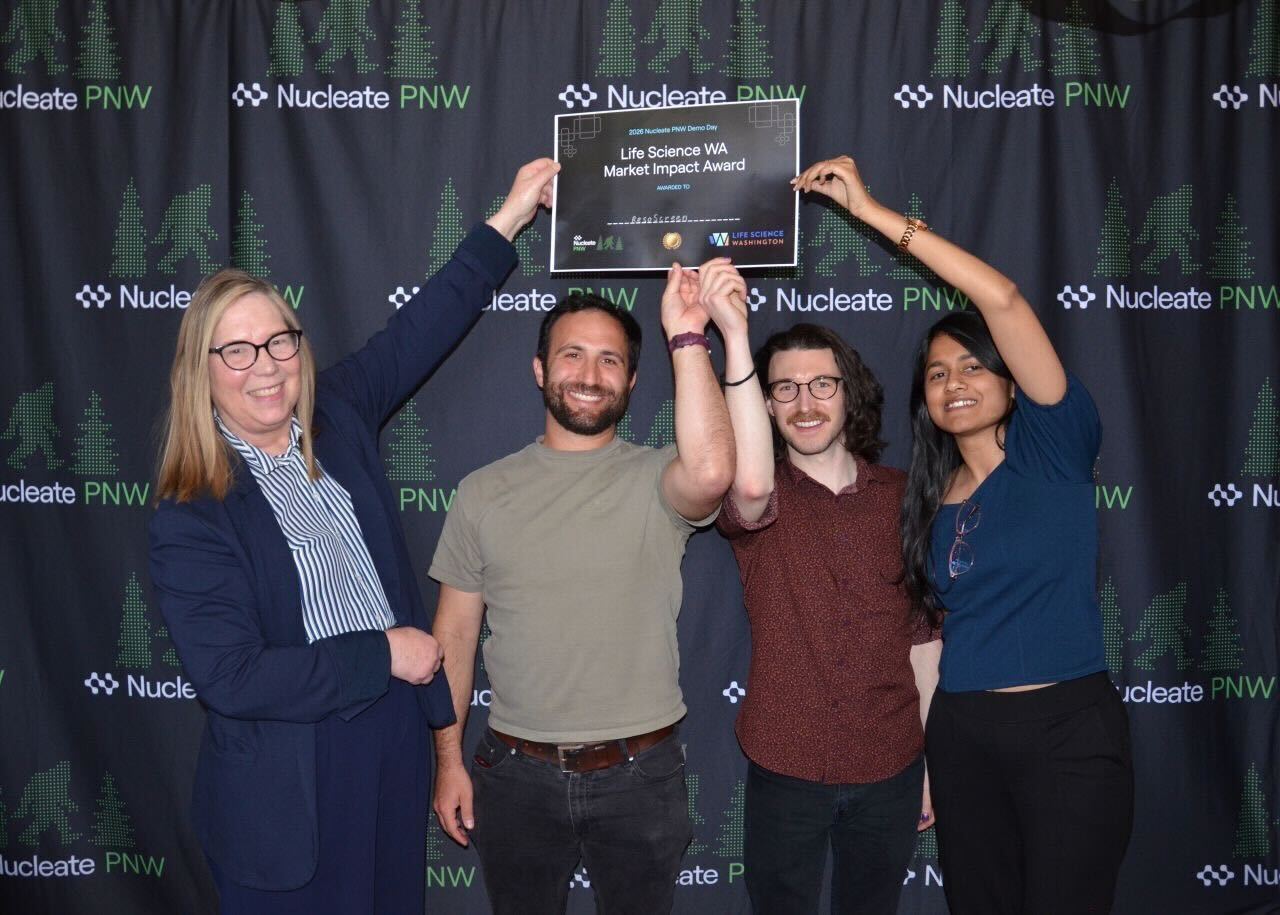 Jason and Rory received the Life Science Washington Market Impact Award for their project ResoScreen at this year’s Nucleate PNW Demo Day
Jason and Rory received the Life Science Washington Market Impact Award for their project ResoScreen at this year’s Nucleate PNW Demo Day -
November 05, 2025
Rory received the 2025-26 Washington Space Grant Dissertation Completion Award! Nice work Rory!
-
June 01, 2024
Congratulations to Keisuke on his new role as an Assistant Professor at Osaka University!
-
August 31, 2023
MISLers are heading to DNA29: Congratulations to professor Georg Seelig for winning the Rozenberg Tulip Award! Also look out for Chandler, Sam, Cady, Carina, and Gwen in Sendai, Japan on Sept 11th-15th
-
March 29, 2023
Congratulations to MISL graduate students Carina Imburgia and Lancelot Wathieu on securing NSF GRFP awards, and MISL undergraduate researcher Kyvalya Reddy on receiving a Mary Gates Research Scholarship!
Research
Nanopore Sensing
We are exploring new and non-traditional ways to use nanopore technology for bespoke molecular sensing, synthetic biology and proteomics.
DNA Tags
Due to its nanoscale size and predictable Watson-Crick base pairing, DNA-based molecular tagging offers an attractive alternative to physical object labeling that is useful when conventional tags, such as QR codes and RFID tags, are unsuitable.
DNA Circuits
We are exploring new approaches to build complex DNA circuits by using automatic microfluidic hardware and array-based DNA synthesis technology.
DNA Storage Architecture and Methods
We are developing methods for reliable and efficient encoding, random access, and decoding of digital data stored in DNA. We are also developing a fully automated whole system architecture.
Microfluidic Automation
We are working on hardware and software to make microfluidics cheaper, more reliable, and easier to use.
DNA Security
We are exploring the intersection of DNA manipulation and security and privacy.
People
Faculty & PIs
Luis Ceze
Jeff Nivala
Karin Strauss
Affiliate Professor
Georg Seelig

Chris Takahashi
Senior Scientists
Lab Manager
Graduate Students

Rory Majule

Samantha Borje

Carina Imburgia

Chandler Petersen
Ashley Stephenson

Daphne Kontogiorgos-Heintz


























